palmerpenguins::penguins %>%
group_by(species, island) %>%
count()
# A tibble: 5 x 3
# Groups: species, island [5]
species island n
<fct> <fct> <int>
1 Adelie Biscoe 44
2 Adelie Dream 56
3 Adelie Torgersen 52
4 Chinstrap Dream 68
5 Gentoo Biscoe 124In today’s #rstats adventure I learnt about tidyr::complete()
r-how-to
Running summarise() or count() and want to keep combinations where there are 0 entries? complete() is your friend!
Quick demo 👇
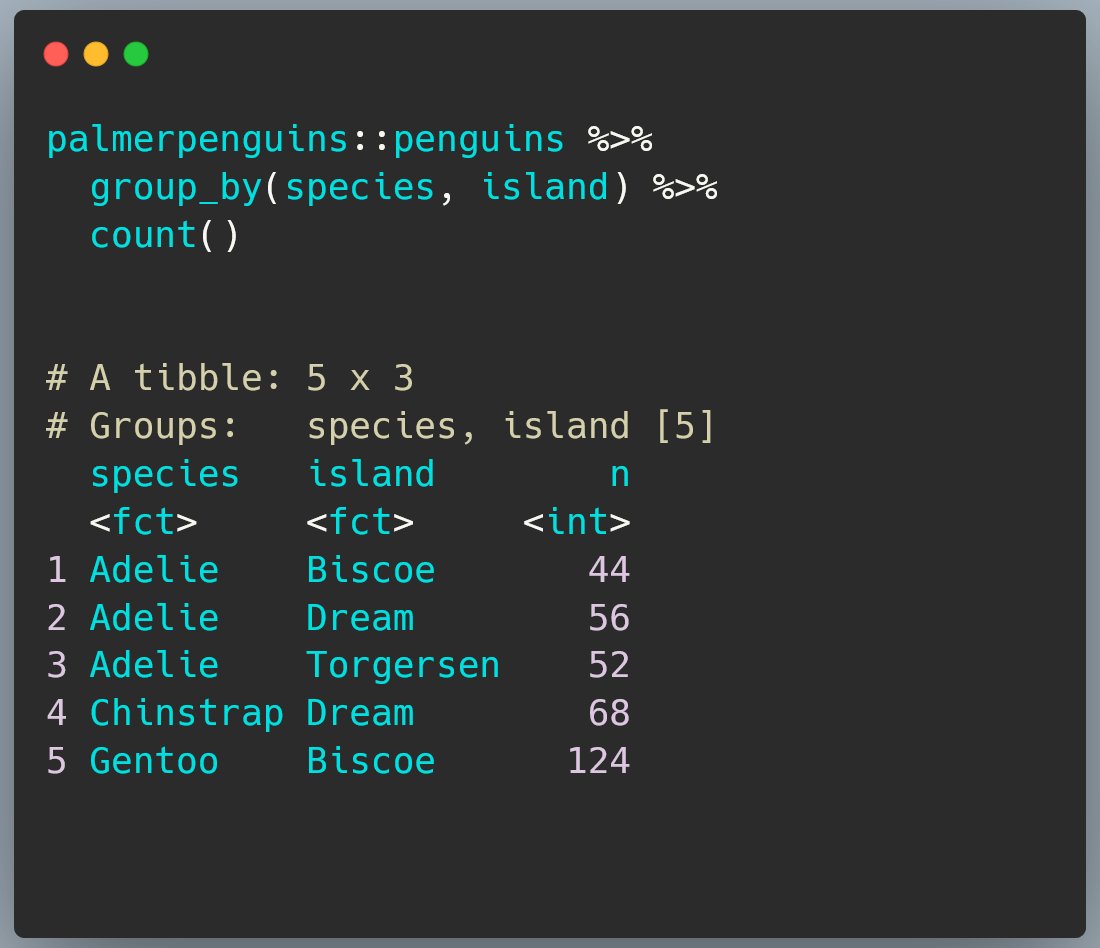
First, the default behaviour.
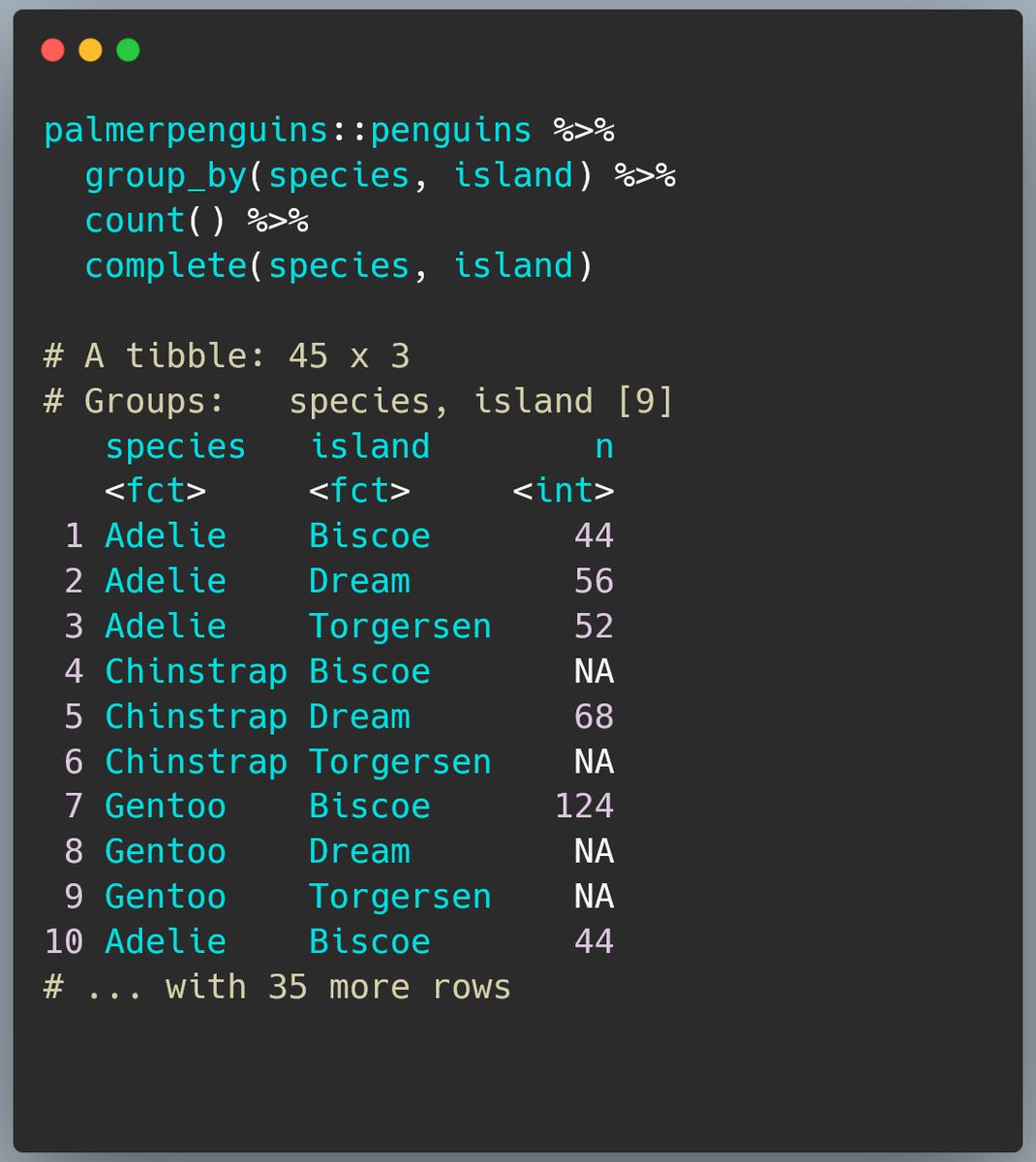
Next, a surprise.
If we just add complete() into the sequence, we end up with 45 rows! There are only 3 penguin species and 3 islands, so we’d only expect 9.
See the note above the table? We have 5 groups in the existing data. 45 = 9 * 5. Aha!
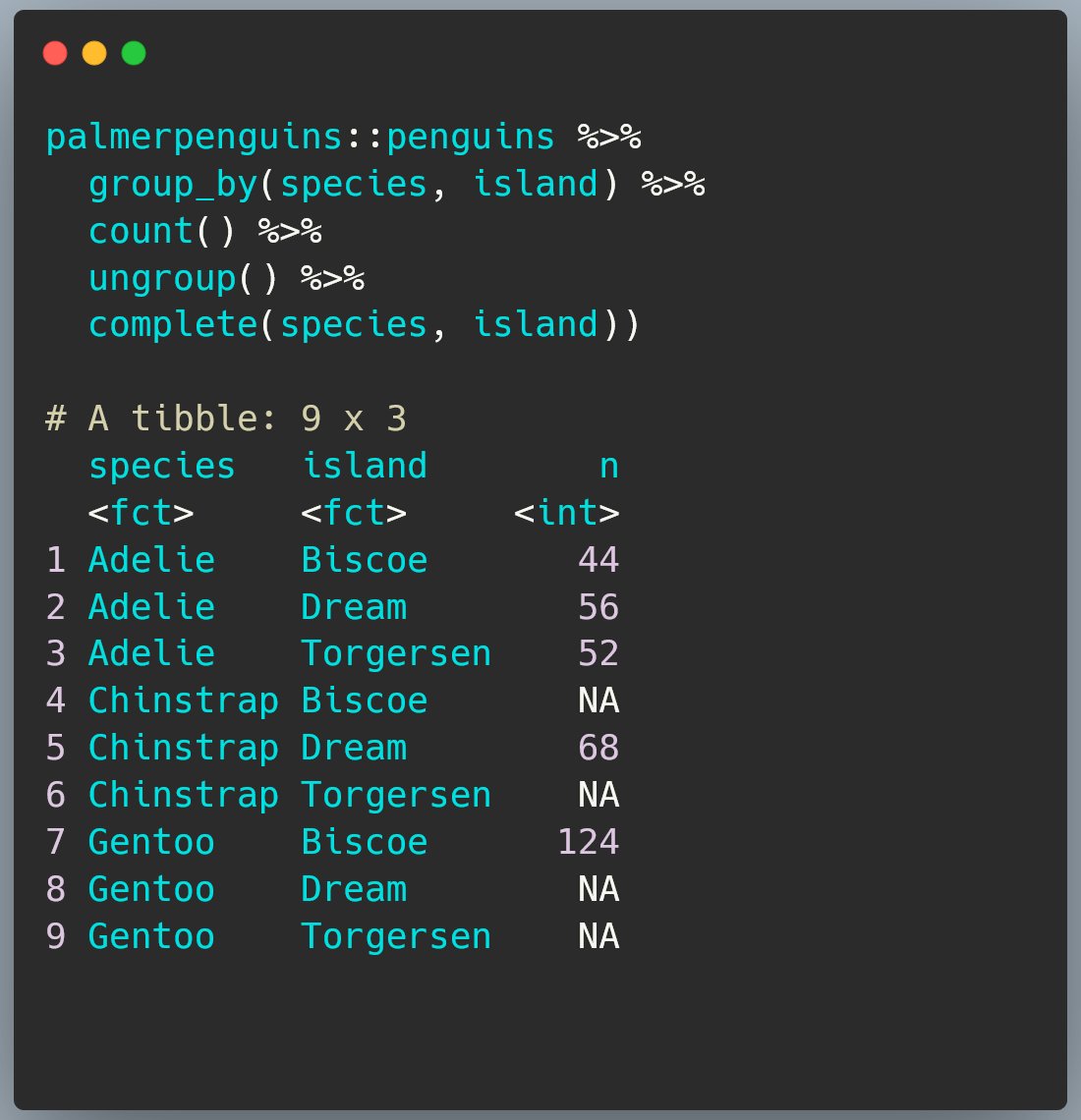
We need to ungroup() before running complete()
Otherwise we’re completing within each current grouping, which isn’t what we want to do.
Tada!
But let’s be more specific
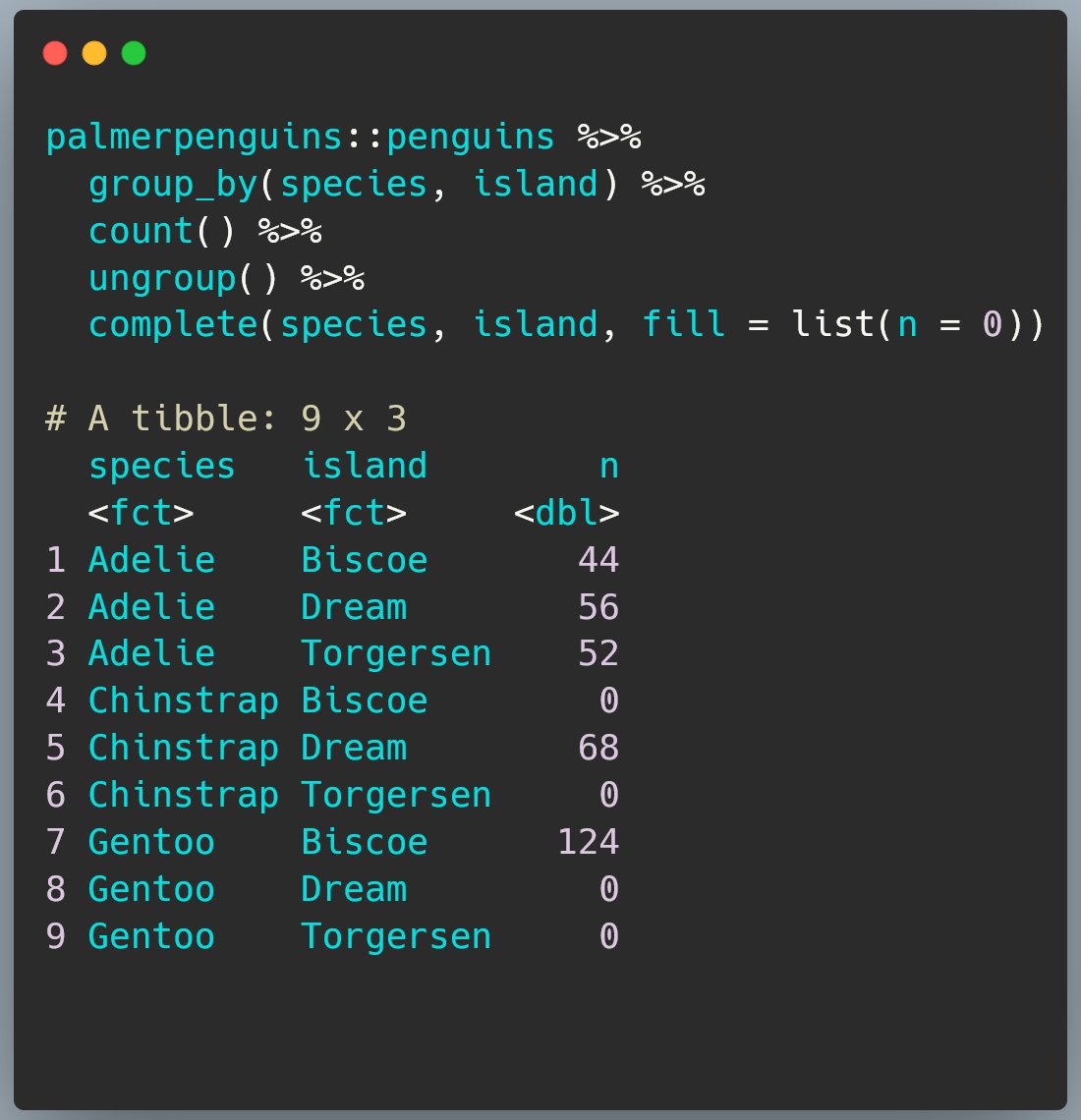
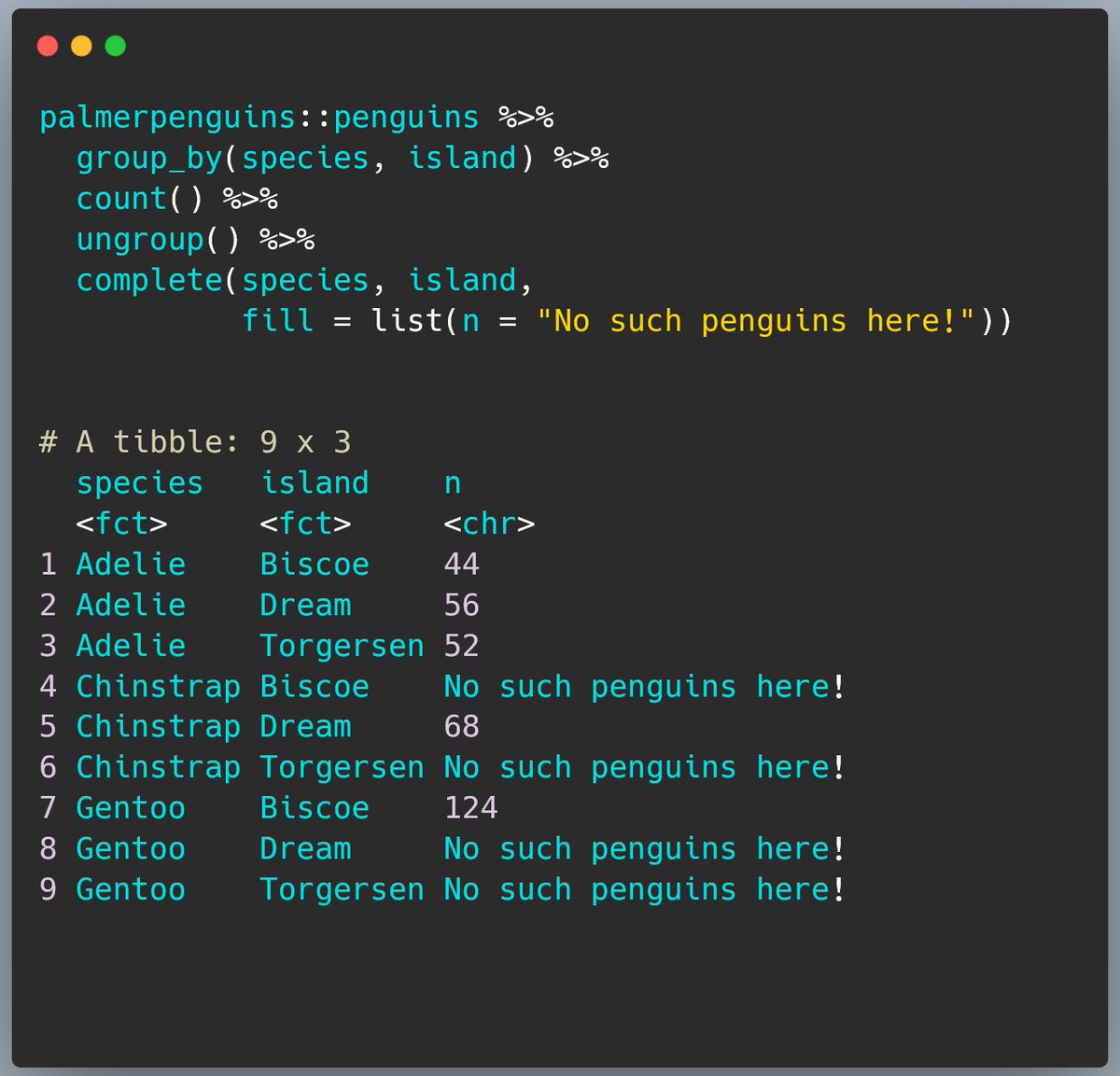
We started this exercise because we wanted to be explicit about there being no penguins of some species on some of the islands so we want to replace the NAs with 0s.
To do that, we need to provide a list of values to swap in for NAs in each column we need to fill. Here that’s n.
A word of warning
If zeros aren’t what you’re after, choose something else. Just beware of that that will do that the class of your column!
Reuse
Citation
For attribution, please cite this work as:
Thompson, Cara. 2022. “In Today’s #Rstats Adventure I Learnt about
Tidyr::complete().” May 26, 2022. https://www.cararthompson.com/posts/2022-05-26-in-todays-rstats-adventure-i/.